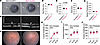Research ArticleGeneticsOphthalmology
Open Access |  10.1172/jci.insight.194681
10.1172/jci.insight.194681
Splicing variants in MYRF cause partial loss of function in the retinal pigment epithelium leading to nanophthalmos
Gabrielle M. Rozumek,1,2 Michelle L. Brinkmeier,3 Bin Guan,4 Su Qing Wang,1 Catherine Tower,3 Nina T. Yang,1 Rachel S. Lim,1 Dejuan Kong,1 Daniel Soden,1 Qitao Zhang,1 John Y.S. Han,1 Jason M.L. Miller,1,5 Lijin Dong,4 D. Ford Hannum,1 Sayoko E. Moroi,5,6 Julia E. Richards,1,7 Robert B. Hufnagel,4,8 and Lev Prasov1,3
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Rozumek, G. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Brinkmeier, M. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by
Guan, B.
in:
PubMed
|
Google Scholar
|

1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Wang, S. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Tower, C. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Yang, N. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Lim, R. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Kong, D. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Soden, D. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Zhang, Q. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Han, J. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Miller, J. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by
Dong, L.
in:
PubMed
|
Google Scholar
|

1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Hannum, D. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Moroi, S. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Richards, J. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Hufnagel, R. in: PubMed | Google Scholar
1Department of Ophthalmology and Visual Sciences,
2Department of Pathology, and
3Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA.
4National Eye Institute, NIH, Bethesda, Maryland, USA.
5Cellular and Molecular Biology Program, University of Michigan, Ann Arbor, Michigan, USA.
6Department of Ophthalmology and Visual Sciences, The Ohio State University, Columbus, Ohio, USA.
7Department of Epidemiology, The University of Michigan, Ann Arbor, Michigan, USA.
8Center for Integrated Healthcare Research, Kaiser Permanente Hawaii, Honolulu, Hawaii, USA.
Authorship note: GMR and MLB contributed equally to this work.
Find articles by Prasov, L. in: PubMed | Google Scholar
Authorship note: GMR and MLB contributed equally to this work.
Published February 26, 2026 - More info
JCI Insight. 2026;11(6):e194681. https://doi.org/10.1172/jci.insight.194681.
© 2026 Rozumek et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: April 22, 2025; Accepted: February 4, 2026
-
Abstract
Improper light focus on the retina, refractive error, is primarily caused by eye size differences and is the leading cause of vision loss worldwide. C-terminal variants in the Myelin regulatory factor (MYRF) gene, a retinal pigment epithelium–derived (RPE-derived) transcription factor, lead to isolated nanophthalmos characterized by a small, though structurally sound eye. However, other MYRF loss-of-function variants cause syndromic disease. To address this discrepancy, in vitro and animal studies were performed on a pathogenic C-terminal variant dG-MYRF (p.Gly1126fs30*, c.3376-1G>A). Human RPE cells or primary RPE transduced with dG-MYRF showed reduced target gene expression, with decreased steady-state levels of the C-terminal cleavage product, but normal cleavage and localization. A homozygous humanized MYRF C-terminal mouse model (MyrfhumdG/humdG) was embryonic lethal by E18.5, while WT (MyrfhumWT/humWT) mice were viable. Single-cell RNA-seq from E17.5 MyrfhumdG/humdG and KO RxCre;Myrffl/fl (E15.5 and P0) mice revealed shared differentially expressed genes, with decreased effect size in the MyrfhumdG/humdG eyes. These findings support dG-MYRF as a hypomorphic allele. Additionally, 2 MYRF splicing variants creating nonfunctional isoforms were found in families with isolated nanophthalmos. Overall, hypomorphic MYRF alleles underlie isolated nanophthalmos, supporting a tissue-specific threshold effect and highlighting unique roles for the MYRF C-terminus in the RPE.
-
Introduction
Myelin regulatory factor (MYRF) is a membrane-associated transcription factor initially identified for its role in oligodendrocyte maturation and myelination (1). It is a type II transmembrane protein that homotrimerizes in the endoplasmic reticulum (ER) lumen and autocleaves to release an N-terminal fragment that translocates to the nucleus and acts as a transcriptional activator (2). More recently, pathogenic variants in MYRF have been associated with an ocular-cardiac-urogenital syndrome featuring congenital heart defects, diaphragmatic hernias, pulmonary hypoplasia, genital abnormalities, and high hyperopia (3–7). MYRF variants have also been described in isolated nanophthalmos, an ocular disease featuring a small but structurally normal eye, with resulting extreme farsightedness (hyperopia). C-terminal frameshift variants have been identified in multiple large pedigrees with predominantly isolated nanophthalmos, suggesting a unique role for the MYRF C-terminus in the eye (8, 9). However, it is not clear why some variants in MYRF result in isolated ocular disease while others produce syndromic phenotypes.
In the eye, MYRF is predominantly and highly expressed in the developing and mature retinal pigment epithelium (RPE) (8, 10). Conditional loss of mouse Myrf in the eye leads to defects in RPE development and retinal degeneration in mice, with perturbations in TGF-β/BMP signaling, pigmentation, cell structure, and cell viability (10). Although the mouse models of Myrf show disparate phenotypes compared with patients, they are invaluable for understanding retinal disease pathogenesis in vivo (8, 11, 12).
Two large nanophthalmos families have been reported with genetic changes that lead to the same C-terminal frameshift mutation in MYRF causing a 31–amino acid extension in the last exon (8, 9). Interestingly, a cluster of patients with pathogenic variants located in the conserved C2 domain manifest with predominantly ocular diseases (Figure 1) (13). While the functional consequences of variants in the DNA binding domain (DBD) and intramolecular chaperone autocleavage (ICA) domain have been well studied (14, 15), there has been little to no progress on understanding the function or importance of the ER-resident, conserved C2 domain.
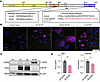 Figure 1
Figure 1Processing of C-terminal MYRF variant in vitro. (A) Diagram of amino acid changes in WT-MYRF, dG-MYRF, and cleavage deficient variant V679A-MYRF. ProRich, proline rich; DBD, DNA binding domain; ICA, intramolecular chaperone auto-processing; TM, transmembrane; C2, C-terminal. (B) Localization of FLAG-tagged MYRF constructs in ARPE-19 cells show normal nuclear localization of dG-MYRF (n = 3). Scale bar: 50 μm. (C) Western blot of transfected ARPE-19 cells shows no change in cleavage of N-terminal fragment (n = 3). (D) qPCR analysis of RNA from ARPE-19 cells transduced with dG-MYRF, compared with WT-MYRF show decreased levels of transcripts for total Myrf and endogenous Tmem98 mRNA (n = 3) by Student’s t test. *P < 0.05, **P < 0.01.
Toward that end, we generated a humanized mouse model of the C-terminal Myrf variant and performed in vitro functional assays in RPE cell lines and primary human RPE cultures to better understand the molecular mechanism by which C-terminal extension alleles cause disease. Furthermore, we identified 2 additional rare splice site variants in MYRF in families with isolated nanophthalmos, supporting this as a more common mechanism for disease pathogenesis. Together, our studies support that partial loss-of-function alleles in MYRF contribute to isolated nanophthalmos and provide mechanistic insights into the role of the MYRF C2 domain.
-
Results
C-terminal MYRF variant is processed normally but shows decreased stability and transcriptional activity. C-terminal MYRF variants and splicing variants are associated with predominantly ocular phenotypes and have familial inheritance, while early truncating and ICA and DBD variants are associated with syndromic phenotypes and predominantly occur de novo (9, 13, 15–17). Given this discrepancy, the C-terminal frameshift alleles could act as dominant negative or hypomorphic alleles. To distinguish these models, we evaluated localization, cleavage, and protein stability of the C-terminal variant in vitro in the ARPE-19 human RPE-like cells. When overexpressed with lentiviral vectors, both WT MYRF and C-terminal frameshift MYRF (dG-MYRF) (p.Gly1126fs30*) showed nuclear localization, while a known cleavage-deficient mutant V679A-MYRF (14) displayed only cytoplasmic localization and was excluded from the nucleus (Figure 1, A and B). Since homotrimerization at the ER lumen is necessary for cleavage of the N-terminal MYRF fragment and its subsequent translocation to the nucleus (1, 2, 18, 19), our results support a model in which dG-MYRF can homotrimerize and cleave.
To further evaluate this, we used Western blotting to systematically define the cleavage dynamics (ratio of full length/cleaved) of the dG-MYRF variant compared with WT-MYRF in ARPE-19 cells. We observed faint bands for full-length FLAG-MYRF (160 kDa) in both WT-MYRF and dG-MYRF and strong transcriptionally active N-MYRF cleavage products (~70 kDa). In contrast, the cleavage-deficient mutant, V679A-MYRF, shows 1 strong full-length MYRF band (Figure 1C). Given normal cleavage and localization, we next evaluated protein stability through computational tools examining degree of disorder Predictor of Natural Disordered Regions (PONDR) and protein folding (AlphaFold2) (20–23). PONDR uses neural networks trained on ordered and disordered regions of short and/or long regions of amino acid sequences from NMR or x-ray crystallography data to predict intrinsic disordered regions. PONDR analysis predicted that dG-MYRF alters the C-terminus from a highly ordered to a highly disordered structure (Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.194681DS1). De novo structural modeling using AlphaFold2 predicted the loss of a β-sheet and alpha-helix structure in the conserved C2 domain of the C-terminal frameshift variant of MYRF (Supplemental Figure 1, B and C). To experimentally validate this, cycloheximide pulse-chase experiments were conducted in ARPE-19 cells transduced with WT-MYRF and dG-MYRF. Protein extracts were analyzed by Western blotting using antibodies to the N-terminus or C-terminus of MYRF. The rates of decay of the C-terminal and N-terminal MYRF cleavage product were unchanged in the dG-MYRF variant following a 24-hour chase (Supplemental Figure 2A). However, steady state levels of C-terminal cleavage product for dG-MYRF were reduced compared with WT (Supplemental Figure 2, B and C). This was unlikely due to reduced epitope detection because the antibody was raised to an epitope present in both forms of MYRF (anti-MYRF393-766AA) (1), and the C2 domain is predicted to fold independently.
To evaluate the functional effect of the dG-MYRF variant on transcriptional activity, we tested the ability of dG-MYRF to autoregulate MYRF transcription and its downstream target TMEM98 using qPCR, as compared with an internal transduction GFP control, present in each of the constructs. ARPE-19 cells transduced with dG-MYRF showed decreased levels of endogenous MYRF and TMEM98 mRNA expression (relative to GFP) as compared with WT-MYRF (Figure 1D).
To evaluate whether RPE differentiation state and polarization alter cleavage dynamics of MYRF, we repeated key ARPE-19 experiments in primary human RPE cultures grown on microporous supports, as described previously (24). We again observed normal localization and cleavage of virally transduced WT-MYRF and dG-MYRF primary RPE cultures (Figure 2, A–D). Additionally, transepithelial electrical resistance (TEER) was measured as a proxy for barrier function and overall cell health since we have previously demonstrated the close relationship between assays for cell toxicity and declines in TEER in primary RPE cultures (25). We observed no difference in TEER in primary RPE transduced with Empty Vector, WT-MYRF, or dG-MYRF 1 week after transduction compared with the same well before transduction (Figure 2E). Overall, the results obtained in the ARPE-19 cell line were consistent with results seen in mature and polarized human primary RPE cultures, demonstrating the dG-MYRF variant does not affect localization, processing, barrier function, or overall cell health but seems to reduce transcriptional activation function of MYRF.
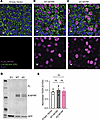 Figure 2
Figure 2Processing and functional studies of C-terminal MYRF variant in human primary RPE. (A–C) Localization of FLAG-tagged MYRF constructs in mature human primary RPE show normal nuclear localization of dG-MYRF. Scale bar: 5 μm. (D) Western blot of transduced human primary RPE cells shows no change in cleavage of N-terminal fragment. (E) Functional assessment of barrier function via TEER (normalized to pretransduction TEER value for each replicate) shows no significant difference in TEER 1 week after transduction with Empty Vector, WT-MYRF, or dG-MYRF by 1-way ANOVA (all experiments are 3 reps from n = 2 donors).
Mouse model of human C-terminal MYRF is prenatal lethal. In vitro analysis of the dG-MYRF variant (Figures 1 and 2) provided insight into the potential molecular mechanism of the patient variant MYRF p.Gly1126fs30*. To understand the pathogenesis of nanophthalmos in patients with the MYRF p.Gly1126fs30* variant, we used CRISPR/Cas9 homology–directed repair to generate a humanized mouse model that mimics the frameshift variant detected in patients, as well as a matched humanized WT control (Figure 3A). In this model, we replaced mouse exon 26 with either the human WT or human C-terminal frameshift exon 27 DNA sequence, including the endogenous-3′UTR sequence, and fused this with mouse exon 25 (Supplemental Figure 3). We previously showed that Myrf is highly expressed in RPE via qPCR of RNA from optic cups and RNAscope in situ hybridization (8, 10). To confirm proper cell specificity and expression of the humanized alleles of Myrf, we used a validated Myrf RNAscope probe to evaluate Myrf expression in MyrfhumdG/humdG and MyrfhumWT/humWT eyes at E15.5. These studies showed consistent Myrf expression pattern and levels among MyrfhumWT/humWT and MyrfhumdG/humdG eyes within the RPE (Figure 3B). To further define the function of the dG allele, we first evaluated survival of MyrfhumdG/humdG embryos. We observed that homozygous MyrfhumdG/humdG embryos were never detected after birth, while MyrfhumWT/humWT mice were viable and fertile. MyrfhumdG/humdG embryos were present in normal Mendelian ratios at E12.5–E17.5 but underrepresented at E18.5 (χ2 prob = 0.015) and at birth (P0) (χ2 prob = 0.042) (Figure 3C).
 Figure 3
Figure 3Homozygous Myrf humdG mice are embryonic lethal. (A) Schematic showing the (humdG) fusion allele amino acid sequence changes in the humanized mouse model. (B) In situ hybridization using RNAscope detected Myrf mRNA in humanized humWT and humdG mouse RPE (n = 4–5 per genotype). Scale bar: 100 μm. (C) Progeny from intercrossing MyrfhumWT/humdG mice exhibit skewed Mendelian ratios at E18.5 (χ2 P = 0.015) and P0 (χ2 P = 0.002). E12.5–E14.5 (n = 35 mice from 4 litters), E15.5 (n = 24 mice from 3 litters), E16.5 (n = 26 mice from 3 litters), E17.5 (n = 44 mice from 5 litters), E18.5 (n = 24 mice from 3 litters), and P0 (n = 56 mice from 5 litters).
Analysis of ocular histology in E16.5 homozygous MyrfhumdG/humdG mice revealed normal pigmentation, which differed from the severe and early-onset depigmentation phenotype we observed in our RxCre;Myrffl/fl mice (Supplemental Figure 4) (8). Tmem98, a direct target of MYRF, is expressed at similar levels in control eyes (MyrfhumWT/humWT) and MyrfhumdG/humdG eyes at the RNA and protein level (Supplemental Figure 5) (8, 26). We cannot rule out the possibility that there are small expression differences not detected by the mRNA and immunostaining assays. However, the RxCre;Myrffl/fl eyes had profoundly reduced expression of Tmem98 (10). The lethality of MyrfhumdG/humdG mice confirm the pathogenicity of the C-terminal Myrf mutant allele, and the lack of obvious ocular defects suggest that it likely functions as a hypomorphic (partial loss-of-function allele) rather than null allele.
MyrfhumdG/+ have normal eye size, retinal and RPE morphology, and retinal function. The RxCre;Myrffl/+ mice had white spots in the retina and RxCre;Myrffl/fl mice had severe retinal degeneration, though no measurable eye size phenotype was observed in either genotype into early adulthood (8). We systematically profiled MyrfhumWT/+ and MyrfhumdG/+ mice over the course of 1 year to determine whether changes ocular dimensions, retinal structure, or function would emerge over time. Noninvasive imaging (SD-OCT and fundus photography) and electroretinography (ERG) were carried out on the same mice over the course of 1-year at 3-month intervals. After 1 year, MyrfhumdG/+ show no significant difference in eye size, axial length, corneal diameter, anterior chamber depth, or retinal thickness and only a modest decline in vitreous chamber depth compared with controls (0.53 ± 0.03 mm versus 0.50 ± 0.01 mm, P = 0.0342) (Figure 4, A–G). Additionally, scotopic and photopic ERGs showed no decrease in visual function, consistent with the SD-OCT findings of normal retinal thickness and morphology (Figure 4, H–J). Fundus photography showed little to no signs of atrophy after 1 year (Figure 4, B and C). To investigate for more subtle changes to RPE morphology similar to those that were evident in RxCre;Myrffl/fl eyes (8), RPE flatmounts were stained with phalloidin, segmented using REShAPE software (27), and evaluated for morphometric features (Supplemental Figure 6). These analyses reveal no differences in the median cell area (P = 0.899), aspect ratio (P = 0.847), or hexagonality (P = 0.7568) of 1-year-old MyrfhumdG/+ eyes compared with MyrfhumWT/+ controls. There was also no change in the median number of neighboring RPE cells between genotypes.
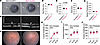 Figure 4
Figure 4Heterozygous MyrfhumdG/+ mice show no gross morphological or retinal eye phenotype after 1 year. (A) Images of enucleated eyes showing similar eye size. Scale bar: 1 mm. (B) Spectral domain optical coherence tomography (SD-OCT) images showing no change in overall thickness or individual retinal layers with white lines representing measurements for vitreous chamber depth (VCD) and retinal thickness. (C) Fundus photographs showing no signs of retinal white spots in MyrfhumdG/+ (pink) compared with controls (black). (D–G) Quantitative assessment of eye size (D), VCD (E), anterior chamber depth (F), and retinal thickness (G) by Student’s t test. (H–J) Retinal function measured by scotopic electroretinogram (ERG) (H and I) and photopic ERG (J) showed no differences (n = 6–8 per genotype). Scale bar: 1 mm.
To evaluate for more subtle transcriptomic changes in MyrfhumdG/+ compared with WT mice, we performed qPCR in P21 adult mouse optic cups including the retina and RPE (n = 4–9 pairs of eyes per genotype). In agreement with the lack of phenotypic changes in these mice, we found no significant differences in gene expression between Myrf+/+ and MyrfhumdG/+ mice in key RPE genes and previously defined MYRF target genes (10) including Mitf, Ermn, Myrf, Sox10, or Tmem98 (Supplemental Figure 7). Together, these results suggest that, at both histologic and the molecular level, MyrfhumdG/+ mice are similar to their matched controls.
To determine the contribution to disease pathogenesis of the Myrf humdG allele, we generated RxCre;MyrfhumWT/humWT, RxCre;MyrfflhumWT, and RxCre;Myrffl/humdG mice for observational studies over a 6-month period to see the effect of the humdG allele on a sensitized background with 1 allele of Myrf missing (Supplemental Figure 8), but the numbers were limited due to poor breeding and survival (n = 4 eyes, 2 mice per genotype). Despite limited sample size, the effect was so severe that it was clear that RxCre;Myrffl/humdG showed signs of RPE degeneration compared with RxCre;MyrfhumWT/humWT and RxCre;Myrffl/humWT controls as early as 2 months and more strikingly by 6 months (Supplemental Figure 8A). In addition, RxCre;Myrffl/humdG mice showed significant reduction in vitreous chamber depth at 4 months (0.4 ± 0.2 mm) versus RxCre;Myrffl/humWT (0.54 ± 0.01 mm, P = 0.0163) and RxCre;MyrfhumWT/humWT (0.525 ± 0.008 mm, P = 0.0106), and 6 months (0.38 ± 0.03 mm) versus RxCre;Myrffl/humWT (0.486 ± 0.005 mm, P = 0.0130) and RxCre;MyrfhumWT/humWT (0.54 ± 0.01 mm, P = 0.0013) (Supplemental Figure 8, B and D). No differences were observed between genotypes in total retinal thickness or overall eye size (Supplemental Figure 8, E and F). In addition to anatomical changes, we observed molecular changes such as mislocalization of a partner protein TMEM98 in 6-month-old RPE flat mounts from RxCre;Myrffl/humdG mice, which was not seen in controls (Supplemental Figure 8C). These data support the Myrf humdG allele as a partial loss-of-function allele.
Single-cell RNA-seq shows altered RPE gene regulatory networks in MyrfhumdG/humdG eyes. To better understand how the C-terminal MYRF variant affects eye development, we performed single-cell RNA-seq (scRNA-seq) on E17.5 optic cups of MyrfhumWT/humWT and MyrfhumdG/humdG mice using the 10X genomics platform (Figure 5A). We chose E17.5, as it is occurs after our RPE pigmentation phenotype (E15.5) is observed in RxCre;Myrffl/fl mice (10) and just before Mendelian ratios become skewed at E18.5. We collected 11,281 and 11,322 cells for MyrfhumWT/humWT and MyrfhumdG/humdG, respectively. The median genes per cell were comparable in WT (2,429) and variant (2,415) samples. Quality control filtering was performed to filter out dead cells, doublets, and poor-quality cells (10). After integration with previously published datasets from conditional KO Myrf eyes (RxCre;Myrffl/fl and Myrffl/fl) at E15.5 and P0, we performed unsupervised clustering and were able to identify 19 clusters, including all the major cell types within the optic cup using previously established cell type–specific markers (Supplemental Figure 9) (10). All clusters were present in both the MyrfhumWT/humWT and MyrfhumdG/humdG mice (Figure 5B), but there was a slight reduction in the RPE cell proportions in MyrfhumdG/humdG mice (5.0%) compared with control (6.4%) (Supplemental Figure 9, C and D).
 Figure 5
Figure 5scRNA-seq reveals molecular changes in RPE of mice carrying C-terminal Myrf Allele (MyrfhumdG/humdG). (A) Schematic depicting optic cups and samples used for scRNA-seq including control and variant mouse optic cups (n = 3 pooled per genotype). (B) UMAPs of control versus Myrf C-terminal variant mouse optic cups demonstrating cell type distributions. (C) Feature plot demonstrating expression of Myrf mRNA in both control and variant mice within the RPE cluster. (D) Volcano plot of differentially expressed genes in MyrfhumdG/humdG mice compared with control (MyrfhumWT/humWT). DEGs were defined by a Bonferroni Padj < 0.1. (E) Top 10 Gene Ontology (GO) Biological Pathway terms for upregulated and downregulated genes.
Myrf expression was detected exclusively in the RPE cluster (Figure 5C), consistent with previous results, leading us to focus our downstream analysis on the RPE cluster. We performed differential gene expression (DEG) analysis between MyrfhumWT/humWT and MyrfhumdG/humdG RPE clusters. We found 735 upregulated and 170 downregulated differentially DEGs with avg_log2FC > ± 0.25 and Padj ≤ 0.1 (Figure 5D), MyrfhumdG/humdG relative to MyrfhumWT/humWT controls. Downregulated genes included extracellular matrix–related (ECM-related) genes Col3a1, Col2a1, Col1a1, and Col1a2 produced by the RPE that play an important role in structure and barrier maintenance (28). Interestingly, some of the most downregulated genes in the variant mice are related to eye size disorders including Serpine3 (avg_log2FC = –1.11, Padj = 2.76E-14), Ankfn1 (avg_log2FC = –1.138, Padj = 2.72 × 10–9), and Kcnq5 (avg_log2FC = –0.81, Padj=6.39 × 10–5) (29–33). Upregulated genes included cell cycle genes Ccnd1 and Cdk8, histone variant components H3f3b (avg_log2FC = 0.71, Padj = 3.69 × 10–34) and H3f3a (avg_log2FC = 0.56, Padj = 3.30 × 10–23), and ubiquitin protein ligase binding genes Tubb5, Ubb (avg_log2FC = 0.59, Padj = 2.32 × 10–27), Tuba1a (avg_log2FC = 0.32, Padj = 0.03). We used overrepresentation analysis (ORA) to identify the top 10 gene ontology terms for Biological Processes and molecular function pathways in humanized variant mice (MyrfhumdG/humdG) and conditional KO (RxCre;Myrffl/fl) relative to their matched controls (MyrfhumWT/humWT and Myrffl/fl). We looked for overlap in top GO terms enriched in MyrfhumdG/humdG and condition KO compared with controls and found shared downregulation of ECM organization (GO:0030198) and extracellular structural organization (GO:0043062). In addition, shared upregulated pathways included structural constituent of ribosome (GO:0003735), structural molecule activity (GO:0005198), rRNA binding (GO:0019843), and translation (GO:0006412) (Figure 5E). Furthermore, GO analysis identified many upregulated pathways in both humanized and KO mice related to metabolism, mitochondrial, and ribosomal function. Thus, the humanized C-terminal variant mutant (MyrfhumdG/humdG) resulted in RPE transcriptome changes similar to the conditional KO (RxCre;Myrffl/fl), including key pathways involved in RPE maintenance and development.
We extended the DEG analysis of the humanized C-terminal variant allele (MyrfhumdG/humdG) and conditional KO (RxCre;Myrffl/fl) by examining individual genes that were affected, including the direction and magnitude of the effect. After integration, we performed DEG analysis on these 2 conditions and looked for genes that had an avg_log2FC ≥ abs(0.25) and Padj ≤ 0.1 in both the C-terminal variant and conditional KO mice compared with controls. We found a total of 55 shared DEGs, with 27 downregulated and 28 upregulated genes respectively (Figure 6A and Supplemental Table 1). Key genes important for RPE development and maintenance were concordant, but the magnitude of the change was decreased in MyrfhumdG/humdG as compared with RxCre;Myrffl/fl mice (Supplemental Figure 10). Some of the concordant downregulated genes and pathways included ECM matrix (Upk3b, Upk1b, Cdh3), Wnt signaling (Tcf4), TGF-β inhibition (Wfkkin2), and Myrf. The concordance of upregulated genes included increases in TGF-β signaling (Id1 and Id3) and glucose metabolism (Gapdh, Ldha, Eno1) (Figure 6B). To validate our findings from that scRNA-seq dataset, we performed qPCR on E16.5 whole eyes (Supplemental Figure 11). We were able to confirm statistically significant increases in expression of Gapdh (1.16 ± 0.26 versus 0.93 ± 0.31 fold change, P = 0.0481) and Ldha (1.33 ± 0.19 versus 0.939 ± 0.19 fold change, P = 0.0267) in MyrfhumdG/humdG mice compared with controls (n = 7–8 pairs of eyes per genotype). Gene set enrichment analysis (GSEA) was performed separately on both MyrfhumdG/humdG and RxCre;Myrffl/fl paired with matched controls, MyrfhumWT/humWT and Myrffl/fl, respectively. GSEA results from MyrfhumdG/humdG RPE emphasized downregulation of ECM and bone development/differentiation pathways, consistent with our prior reports, which showed similar gene enrichment in RxCre;Myrffl/fl RPE. Notably, pathways related to eye and sensory organ development were absent in the top GSEA results for MyrfhumdG/humdG, while they dominated the top results for RxCre;Myrffl/fl (Figure 6, C and D).
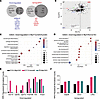 Figure 6
Figure 6MyrfhumdG C-terminal variant acts as a hypomorphic allele. (A) Comparison of differentially expressed genes in RPE cluster of E17.5 humanized C-terminal variant (MyrfhumdG/humdG) versus conditional KO (RxCre;Myrffl/fl) mice with key shared genes highlighted (n = 3 pooled per genotype). (B) Plot of MyrfhumdG/humdG versus RxCre;Myrffl/fl fold change in gene expression for genes expressed in both datasets (log2FC > 0.25, Bonferroni Padj < 0.1) showing strong concordance of up- and downregulated differentially expressed genes between the humanized allele and conditional KO. (C and D) Gene set enrichment analysis (GSEA) showing top 10 downregulated pathways in MyrfhumdG/humdG versus RxCre;Myrffl/fl mice. (E and F) Pseudobulk analysis of RPE cluster demonstrating magnitude of downregulation compared with respective control (E) is lower in the C-terminal variant (pink) than the conditional knock out at E15.5 (black) or P0 time points (blue), while magnitude of upregulation is similar across groups (F).
Finally, to assess the retained function of the humdG allele relative to conditional KO, we performed pseudobulk analysis on the RPE clusters and compared expression of select downregulated (Figure 6E) or upregulated (Figure 6F) genes between MyrfhumdG/humdG (E17.5) and RxCre;Myrffl/fl (E15.5 and P0). We observed that the MyrfhumdG/humdG RPE retained 46%–90% expression across various downregulated genes as compared with P0 RxCre;Myrffl/fl mice, which only retains 1%–66% expression as compared with their respective controls. Interestingly, for select upregulated genes, we observed no differences in expression between the genotypes, suggesting these are less likely to be direct targets and may be buffered. Together, these data support shared pathogenic mechanisms between the KO model and the C-terminal variant, stemming from hypomorphic MYRF effects at the level of RPE.
Deep intronic variants in MYRF in nanophthalmos patients alter splicing. As our in vitro and transcriptomic findings suggest a hypomorphic effect for the dG-MYRF variant, we reasoned that other MYRF splicing variants may also cause ocular only phenotypes, either by affecting only the C-terminal domain exons or by causing a partial reduction in functional RNA or protein products. To validate this hypothesis, panel-based or whole genome sequencing (WGS) was carried out on a cohort of 53 families with nanophthalmos/high hyperopia that did not have identified genetic diagnoses (34). From this cohort, 2 families were identified with deep intronic variants in MYRF (NM_001127392.3 MYRF c.460+167G>A in P01965 from Family 1 and c.3194+122A>G in P04818 and P04825 from Family 2) (Figure 7, A and B). No other rare deleterious genetic variants in nanophthalmos-associated genes (MYRF, PRSS56, MFRP, CRB1, BEST1) were identified in either family.
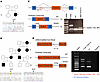 Figure 7
Figure 7Deep intronic variants in MYRF alter splicing of mRNA transcript. (A and B) Two families were discovered with intronic variants in MYRF. (C) Schematic of the minigene RNA splicing assay and predicted products. Minigene for variant c.460+167G>A (A) show altered splicing of MYRF at the mRNA level for 39% of transcripts from sample from Family 1, suggesting a ~19% usage overall (n = 18 colonies). (D) cDNA and predicted splicing of c.3194+122A>G (B) variant amplified from MYRF RNA in patient blood from Family 2. PCR amplification of cDNA shows altered splicing at the mRNA level for 15% of transcripts (30% for the allele) (n = 20 colonies).
Clinical examination of family members revealed very short axial lengths in both families and classic features of nanophthalmos including choroidal folds and retinal detachment (Supplemental Figure 9 and Supplemental Table 2). Proband P01965 demonstrated no family history of nanophthalmos (Figure 7A), while Family 2 exhibited a pattern of autosomal dominant inheritance, with demonstrated segregation in 2 generations. No systemic features of MYRF cardiac-urogenital syndrome were noted in any of the affected individuals or other family members.
The high spliceAI (35) scores for the variants in Family 1 and 2 (c.460+167G>A, score = 0.992 and c.3194+122A>G, score = 0.8, respectively) indicate that the variants have a high probability of altering splicing (Supplemental Table 3). The first variant was predicted to cause gain of a cryptic acceptor site (c.460+167G>A), leading to pseudoexon insertion of 91 bp (Figure 7C) and a frameshift causing either early protein truncation or RNA degradation via nonsense mediated decay. An exon trap minigene splicing assay confirmed that the c.460+167G>A variant altered splicing compared with WT MYRF, producing a 374 bp fragment corresponding to the predicted pseudoexon inclusion in addition to the normal splicing product (283 bp WT exon 5 amplicon) (Figure 7C). To quantify the fraction of transcripts that have aberrant splicing, PCR fragments were TA cloned, and individual colonies were sequenced (n = 18). Seven of 18 (38.9%) contained RNA produced from the cryptic splice site usage, while the remaining used the normal splice acceptor.
For Family 2, we confirmed cosegregation of the variant in 2 generations, yet we did not detect pseudoexon inclusion using the exon trap vector system (Supplemental Figure 13, A–C). The variant may have a cell type–specific or contextual effect on splicing not detected by our assay or affect MYRF function through a different mechanism. To support this possibility, we extracted RNA from freshly collected patient blood and performed RT-PCR to look for evidence of altered transcript splicing. We observed an additional ~300 bp band matching the expected size of an alternatively spliced transcript for this variant (Figure 7D). To quantify the fraction of transcripts that have aberrant splicing, PCR fragments were TA cloned, and individual colonies were sequenced, showing 3 of 20 colonies (15%) using the cryptic splice site usage, and suggesting ~30% usage of the cryptic splice site from that allele overall (Figure 7D and Supplemental Figure 13D). The predicted protein product based on cDNA sequencing would cause a frameshift leading to early truncation after exon 24 (Supplemental Figure 13E).
Together, these data suggest that intronic splice variants in MYRF may be another mechanism leading to the pathogenesis of nanophthalmos. Isolated ocular phenotypes may be driven by partial loss-of-function alleles with incomplete alternative splice site compensation or altered function of the alternative transcript’s protein product.
-
Discussion
Though originally identified as a master regulator of oligodendrocyte maturation, growing evidence supports a role for MYRF in regulating RPE development, differentiation, and maturation (8, 10). Our previous studies underscore the importance of MYRF in the developing RPE, showing that conditional loss of Myrf in mice can be disastrous for RPE development and lead to retinal degeneration (8). Interestingly, there are a cluster of variants located in the conserved C2 domain that manifest as isolated ocular diseases (13). Combined with other partial loss-of-function alleles, including splicing variants we further identify in this study, our data support that the C2 domain variants and other hypomorphic alleles may manifest as isolated ocular disease (Figure 8).
 Figure 8
Figure 8Working model of MYRF splice variant functional and phenotypic effects contributing to isolated ocular and syndromic disease pathogenesis. WT MYRF retains transcription factor function resulting in no phenotypic effects. A C-terminal frameshift splice variant results in a 31 amino acid frameshift leading to decreased C-terminus stability and transcriptional activation, leading to reduced transcription factor function and isolated ocular disease. Intronic splicing variants lead to a frameshift that results in early truncation or nonsense mediated RNA decay. These splice sites are not always used, leading to modest decrease in MYRF transcription factor function and isolated ocular disease. In contrast, complete loss of function alleles either lead to early truncation/nonsense-mediated decay or alter critical functional domains (i.e., DNA binding, cleavage) and lead to severe, syndromic symptoms.
C-terminal MYRF variant acts as a loss-of-function allele. We show 4 lines of evidence supporting that the C-terminal MYRF variant is a partial loss-of-function allele. First, embryonic lethality is observed in mice harboring a homozygous C-terminal truncating allele in Myrf, in line with the autosomal dominant inheritance pattern and lack of homozygous mutant individuals in humans. Embryonic lethality is observed later in gestation than constitutional KO of Myrf, with skewed Mendelian ratios starting at E17.5, rather than lethality during cardiac development midgestation (36). Second, there are many shared disrupted transcriptional targets and pathways, in our MyrfhumdG/humdG mouse model and the RxCre;Myrffl/fl conditional KO mice, including ECM, TGF-β signaling, Wnt signaling, structural regulation, and RPE maturation pathways, and regulation of Myrf itself, arguing against neomorphic function.
Third, transcriptional and phenotypic effects in MyrfhumdG/humdG were milder than in comparable RxCre;Myrffl/fl mice. MyrfhumdG/humdG had normal-appearing RPE pigmentation, modest reduction in Tmem98 expression, and lower magnitude of DEGs among shared pathways with the RxCre;Myrffl/fl eyes. Additionally, no phenotypic changes in the heterozygous MyrfhumdG/+ eyes were apparent over 1 year. The direct comparison of scRNA-seq datasets with the conditional KO suggests that the C-terminal MYRF variant allele retains some partial function. Although molecular changes are picked up on scRNA-seq data, the phenotypic changes in the MyrfhumdG/humdG are more modest in comparison with the RxCre;Myrffl/fl mice. For example, genes in the pigmentation pathway are present in the downregulated DEGs of MyrfhumdG/humdG such as Oca2 at E17.5, but these mice show no signs of depigmentation at embryonic stages. Fourth, the C-terminal frameshift MYRF allele exhibits only subtle processing defects, with decreased steady state stability and transcriptional activity in vitro, but no differences in protein, localization, or cleavage in the context of an overexpression assay.
A previous study developed a mouse model to investigate a similar variant, MYRF c.3260delG/p.(Gly1087Valfs*151) mutation, which corresponds to the human MYRF c.3377delG/p.(Gly1126Valsf*31) mutation (9). This mutation causes the same C-terminal frameshift in humans as the c.3776-1G>A discovered by our lab. While there was support for loss-of-function for this allele, with lower mRNA and protein expression of MYRF (9), the mouse allele did not mimic the human allele, given the difference in the frameshift amino acid sequence and the-3′UTR sequence. Furthermore, comparison of embryonic lethality of the Gly1087Valfs*151 variant allele to our current study is not possible, as the prior study only evaluated heterozygous mice. Therefore, our humanized mouse has more fidelity to model the human disease by replacing the last exon with human WT or variant DNA sequence and recapitulates the 31 amino acid frameshift in patients. Some discrepancies remain between our model and the previously published one (9), as we did not find evidence to support changes in thickness of GCL or INL layers (measured by SD-OCT), phenotypic/morphological changes in heterozygous mice, or signs of primary angle closure glaucoma. These may be related to the different nature of the allele, CRISPR off-target effects, or strain/background specific effects. Our humanized mice and WT mice exhibit mRNA expression exclusively within the RPE cluster; therefore, antibody detection of MYRF in other eye structures may represent cross-reactivity or background (8, 10). Our data, together with these previous models, support the assertion that C-terminal Myrf variant is acting as a partial loss-of-function allele. However, future studies are necessary to completely rule out the possibility of the C-terminal variant acting via dominant negative interactions with the WT MYRF, as these mechanisms could not be assessed easily in our animal model or overexpression cell culture models.
Eye size and the dG-MYRF mouse model. In both our animal model of the human allele and our prior developmental KO model (8, 10), axial lengths were comparable with those of the control, suggesting either that the mouse may not recapitulate all the features of human nanophthalmos or that more efficient KO or longer follow-up is necessary to see this phenotype. The 2 mouse models of recessive nanophthalmos genes, MFRP and PRSS56, with the most severe axial length reductions in humans (34, 37, 38) have produced only modest reductions in axial length (<0.15–0.2 mm) (39–41). Mouse models in other nanophthalmos-associated genes, including BEST1, CRB1, and TMEM98 do not show reductions in axial length (42–46) but do show various degrees of reduced retinal function and/or retinal degeneration, consistent with our models of Myrf (40). Furthermore, all 3 of the RPE-derived genes (Best1, Tmem98, and Myrf) are expressed in adult RPE, suggesting a potential maintenance role in RPE. Complex molecular signaling between the RPE and retina may lead to compensation in ocular dimensions, leading to no observable changes in eye size. For example, upregulation of Prss56 is observed in Mfrp rd6-KO mice and AAV2 mediated overexpression of Prss56 alone in adult mouse eyes can increase axial length (41). Thus, Prss56 upregulation may temper any reduction in eye size in the Mfrp-KO mouse. Interestingly, in our cohort of RxCre;Myrffl/humdG mice we saw limited effects on retinal thickness but observed a significant decrease in VCD after 4 and 6 months compared with either RxCre;Myrffl/humWT or RxCre;MyrfhumWT/humWT. Changes in VCD are correlated to eye size and refractive error in humans and mice (47–50). Measurements of overall eye size may be complicated by the technique and variability, and future studies will be necessary to fully evaluate the contribution of MYRF to eye size phenotypes in mice. Finally, our data suggest that eye size defects may precede retinal degeneration, which was seen more robustly in our RxCre;Myrffl/fl–conditional KO model. Further work to explore the compensatory mechanisms buffering eye growth will be informative to address the discrepancies between human disease and mouse models.
Consequences of functional loss of C-terminal domain. Our results show that the C-terminal MYRF variant can cleave and localize properly to the nucleus like the WT protein. This is consistent with previous work that demonstrated a truncated version of MYRF lacking the C-terminal domain (MYRF-1:756) maintains proper homotrimerization and processing in oligodendrocytes (2). Previous work has also shown that homotrimerization is necessary for MYRF self-cleavage (1). This implies that the C-terminal variant is also able to form the homotrimer structure necessary to facilitate cleavage of MYRF. As the C-terminal MYRF variant homotrimerizes and cleaves, we might expect it to be capable of activating transcription of its downstream targets. However, our scRNA-seq and qPCR data show downregulation of key target genes and pathways involved in RPE development including the Wnt signaling, TGF-β signaling, pigmentation, cytoskeletal organization, and epithelial cell development. The changes in these pathways are consistent with many changes detected in our conditional KO model. For example, top downregulated hits in MYRF C-terminal variant RPE included known eye size disorder genes. Serpine3 encodes a serine protease inhibitor, which has been shown to be significantly upregulated in induced myopia models in zebrafish and chicks (30, 31). In contrast, it is significantly reduced in our model, suggesting that it may play a bidirectional role in eye size regulation like Prss56. Additionally, other top downregulated genes Ankfn1 and Kcnq5 are known GWAS hits for refractive error and/or myopia (51–53). Downregulated Ankfn1 mRNA expression has recently been discovered as a defining feature of healthy versus dedifferentiated RPE in mouse models that are aged or exposed to cigarette smoke (29). Kcnq5 encodes a voltage-gated potassium channel expressed in the RPE (human, primate, and bovine) and involved in active transport of K+ from the retina to the choroid (32, 33). These data suggest that the C-terminal variant is likely influencing expression of genes involved in eye size regulation in the RPE and demonstrates how leveraging scRNA-seq technology can help to uncover early changes in gene expression, which are challenging to detect via bulk RNA-seq or qPCR in complex tissues like the developing embryonic eye.
Interestingly, although dG-MYRF maintains normal protein processing, at steady state, the full-length and C-terminal fragments are less stable. It is possible that the instability observed at steady state levels may be a driving factor behind decreased transcriptional activation of downstream target genes. Furthermore, with in vitro overexpression, we see loss of MYRF autoregulation and inability to upregulate the TMEM98 downstream target in the dG-MYRF mutant. These results are consistent with other studies supporting a role for the C-terminus of MYRF in regulation of its function, as removal of the ER-resident portion in vitro results in decreased levels of target gene transcription (2, 54). Furthermore, overexpression of a mouse Myrf plasmid missing the ER portion results in decreased expression of genetic markers of differentiation in maturing oligodendrocytes (2). Together, these data suggest that the C-terminal MYRF variant and removal of the ER portion may decrease target gene expression, even though the protein appears to maintain normal cleavage and localization dynamics in cell culture, suggesting that other important protein-protein interactions may be at play.
Presence of splice-altering variants in nanophthalmos. In addition to variants in well-characterized domains such as the DBD and the ICA cleavage domain, which fundamentally alter MYRF’s function to act as a transcription factor, we found predicted splice variants in MYRF that are associated with nanophthalmos. We discovered what we believe to be two novel, deep intronic variants in MYRF that are predicted to alter splicing; one of these was confirmed by a minigene splicing assay and the other by directly assessing patient blood RNA for alternatively spliced transcripts. The splicing variant from Family 1 is expected to result in an early frameshift and produce a nonfunctional protein product. The minigene splicing assay suggests that pseudoexon inclusion occurs in ~40% of the transcripts from that allele. The splicing variant from Family 2 is expected to create a stop codon after exon 24 and predicted to create a truncated protein product with reduced function. Splicing from patient blood suggests that the pseudoexon inclusion occurs in ~30% of the transcripts from that allele. These data, together with our animal and cell culture evidence from the C-terminal frameshift variant, support a tissue-specific, dosage-sensitivity threshold for MYRF, whereby partial loss-of-function variants may lead to ocular disease only, while complete loss-of-function or dominant negative alleles may lead to syndromic disease. Dosage sensitivity of MYRF is supported by previous studies showing intrafamilial phenotypic variability (8, 17, 55). Indeed, most individuals with cardiac-urogenital syndrome that are evaluated for ocular phenotypes do have high hyperopia (3). Since WGS is applied more often in clinical settings, deep intronic variants in MYRF and other nanophthalmos-associated genes may be found. Furthermore, WGS may improve our ability to capture DNA variants in highly conserved noncoding regions such as promoters or enhancers that may be regulated by MYRF. One limitation is that MYRF variants are a rare cause of nanophthalmos and, therefore, we may not observe the exact same variants in multiple families. However, we expect WGS to reveal similar splicing alternations that lead to production of nonfunctional transcripts. Additionally, future studies could benefit from implementation of long-read sequencing to resolve additional splice isoforms (at the RNA level) or any changes in RNA modifications in MYRF (56) and allow for haplotype resolution and assessment of epigenetic marks (56, 57) at the DNA level.
Conclusions. In this study, we have investigated mechanisms by which MYRF alleles may cause isolated ocular disease. Our results emphasize the role of the protein’s C2 domain in regulating of the activity of MYRF in the RPE, as its alteration leads to decreased steady-state stability, transcriptome-wide dysregulation, and downregulation of pathways important for RPE development. We also demonstrate that the C-terminus of MYRF is important and necessary to produce viable offspring in mice. These results build upon the previously existing limited knowledge of the function and importance of the conserved C-terminal domain of MYRF and offer steppingstones to further understand how C-terminal variants of MYRF contribute to disease pathogenesis. Furthermore, our identification of variant alleles that alter splicing, while maintaining some appropriate splicing, supports the conclusion that the eye is particularly sensitive to reduced MYRF function.
-
Methods
Sex as a biological variable. Our study examined male and female animals, and similar findings are reported for both sexes.
Plasmid generation and lentiviral transduction of ARPE-19 cells. Human FLAG-tagged WT-MYRF DNA constructs were generated (Twist Bioscience, South San Francisco, CA) in a pLentiLox-IRES-GFP backbone vector (58). The C-terminal MYRF mutant and the V679A cleavage deficient were generated by site-directed mutagenesis of the pLentiLox-FLAG-MYRF-IRES-GFP plasmid using partially overlapping primers (Forward: 5′ GCACTGCTGGTCAGGCCAACTGCAGTTCAGAGG-3′, Reverse: 5′-TGGCCTGACCAGCAGTGCCACCCGAAAGTGGTA-3′) and the Quik Change II Kit (Agilent Technologies, Santa Clara, CA, #200517) (Supplemental Table 5) and verified by junctional Sanger sequencing, and whole plasmid sequencing (Plasmidsaurus, South San Francisco, CA). Lentiviral constructs were packaged by the University of Michigan Vector Core (10X concentrated virus) using standard methods. Viral titers were assessed by flow cytometry for GFP on subconfluent ARPE-19 cells with assistance from the University of Michigan Flow Cytometry Core.
ARPE-19 cell culture. ARPE-19 cells (CRL-2302,American Type Culture Collection Manassas, VA) were grown according to established protocols (59, 60) on DMEM/F-12 HEPES media (Thermo Fisher Scientific, Waltham, MA, #11330032) supplemented with 10% fetal bovine serum (FBS) (Corning, Glendale, AZ, #35-010-CV), 10 mM nicotinamide (Sigma Aldrich, Burlington, MA, #N0636), and 1% penicillin/streptomycin (Gibco, Waltham, MA, #15140-122), and passaged 1:4 on P100 plates.
Human primary RPE cultures and TEER. Primary human RPE cultures (hRPE) were grown according to established protocols (24, 61–64). Cells for localization studies were seeded on passage 1 and grown on microporous inserts (Transwells [Corning, Glendale, AZ, USA], product 3470) for at least 2 months. All primary cells were genotyped for Age Related Macular Degeneration risk SNPs: CFH (Y402H, rs1061170), C3 (R102G, rs2230199), C2 (rs9332739), and HTRA1/ARMS2 (rs3750846) using PCR amplification with primers in Supplemental Table 4 followed by ExoSAP-IT PCR Product Cleanup (Thermo Fisher Scientific, Waltham, MA, #78200.200.UL) and Sanger sequencing (Eurofin, Louisville, KY). No high-risk SNPs were detected.
Protein localization studies. ARPE-19 cells were seeded at 40,000 cells/well on Nunc Lab-Tek II 4-well chamber slides (Thermo Fisher Scientific, Waltham, MA). After 24 hours, cells were transduced with 1 × 10–2 dilution of WT-MYRF, C-terminal mutant MYRF, or V679A (10X concentrated) vectors in the presence of 100 mg/mL LentiBoost (Mayflower Biosciences, St. Louis, MO, #SB-P-LV-101-11). Cells were fixed with 4% PFA (paraformaldehyde) in 1× PBS (phosphate buffered saline) for 10 min, incubated with rabbit anti-FLAG 1:1,000 (Cell Signaling Technology, Danvers, MA, #14793) and mouse anti-Calnexin 1:200 (Sigma Aldrich, Burlington, MA, #C4731) primary antibodies overnight at 4°C followed by incubation with anti-goat rabbit Alexa Fluor 488 1:500 (Jackson ImmunoResearch Labs, WestGrove, PA, #111-545-144) and goat anti-mouse Alexa Fluor 555 1:1,000 (Invitrogen, Carlsbad, CA, A-21422) secondary antibodies for 2 hours at room temperature (RT). Cells were washed 3× 5 min with 1×PBS,stained with DAPI (Thermo Fisher Scientific, Waltham,MA), mounted with gold anti-fade media (Thermo Fisher Scientific, Waltham, MA, P36930), and imaged at 63X using Leica SP8 confocal microscope (Leica, Wetzlar, Germany).
Confluent and mature hRPE were transduced at an MOI of 5 with either an Empty, WT-MYRF, or C-terminal mutant MYRF (10X concentrated) vectors in the presence of 100 mg/mL LentiBoost. After 1 week, cells were fixed with a 4% PFA + 4% sucrose solution for 15 min. Pigment was removed from cells by photochemical bleaching in a bleaching solution (250 μL deionized formamide, 4 mL water, 250 μL of 20× Saline Sodium Citrate (SSC) (1× final), 817 μL 30% hydrogen peroxide) for 15 minutes under a stadium light (Yanycn, power: 400W LED, color: 5000k daylight white light), and then washed 5 times with 1×PBS. The reaction was then quenched with 50 mM NH4Cl for 10 minutes and washed 2–3 times with 1×PBS (62).
After bleaching, fixed cells were permeabilized with 0.25% TritonX-100 in 1×PBS for 10 minutes, blocked for 1 hour at RT, and then incubated with rabbit anti-FLAG 1:1,000 or mouse anti-Calnexin 1:200 primary antibodies for 1 hour at RT. Fixed cells were then incubated with anti-goat rabbit Alexa Fluor 647 1:1,000 (Invitrogen, Carlsbad, CA, #A-21446) and goat anti-mouse Alexa Fluor 594 1:1,000 (Invitrogen, Carlsbad, CA, #A-11005) secondary antibodies for 1 hours at RT. Z stacks were obtained using the Leica STELLARIS 8 FALCON Confocal (Leica, Wetzlar, Germany). Microscope and image analysis was performed using IMARIS 10.2 software (Oxford Instruments, Abingdon, England).
Protein stability and cleavage studies. ARPE-19 cells were seeded at 40,000 cells in 24-well tissue culture plates. After 24 hours, subconfluent cells were transduced with lentiviral vectors carrying WT-MYRF, C-terminal mutant MYRF, or V679A (MOI 3 from 10X concentrated stocks) vectors in the presence of 1 mg/mL LentiBoost. Cells were treated with 300 μg/mL cycloheximide (Sigma Aldrich, #C4859-1ML) and lysates collected in Pierce RIPA Buffer (Thermo Fisher Scientific, #89900) following 2 cell rinses in PBS at the following time points: 0, 12,16, 20, 24 hours. Protein concentration of cell lysates was quantified using Pierce BCA Protein Assay Kit (Thermo, #2325). Samples were denatured using 4× Laemmli Sample Buffer (Bio-Rad Laboratories, Hercules, CA) supplemented with 2-Mercaptoethanol and boiled at 95°C for 5 min. Upon SDS-page, samples were transferred onto Immobilon-FL PVDF membrane (Millipore Sigma, Burlington, MA) and probed chicken anti-GFP 1:5,000 (Abcam, Cambridge, England, #13970) or rabbit anti-GAPDH 1:1,000 (Millipore Sigma, #ABS16), as well as rabbit anti-FLAG 1:1,000 or mouse anti-MYRF393-766 (1) (gifted from Ben Emery, Oregon Health & Science University, Oregon, USA) primary antibodies. IRDye 800CW goat anti-rabbit IgG 1:20,000 (LICOR, Lincoln, NE, 926-32211) and IRDye 680RD donkey anti-chicken IgG 1:20,000 (LICOR, Lincoln, NE, #925-68075) were used as secondary antibodies. Blots were imaged using the Odyssey CLx and quantified using ImageStudio (LICOR).
qPCR. For ARPE-19 experiments, cells were seeded at 50,000 cells/well on 24-well plates. After 24 hours, cells were transduced with 1 × 10–2 dilution of WT-MYRF, C-terminal mutant MYRF, or V679A (10X concentrated) vectors in the presence of 100 mg/mL LentiBoost. After 72 hours, RNA was isolated from cells using the RNAqueous-Micro Total RNA Isolation Kit (Invitrogen, Carlsbad, CA) according to the manufacturer’s protocol. cDNA was generated using the SuperScript III system (Thermo Fisher Scientific, Waltham, MA) following the recommended protocol. qPCR was performed on the Applied Biosystems 7500 Real Time PCR System (Applied Biosystems, Foster City, CA) using the TaqMan Assay system with inventoried probes for MYRF, TMEM98, B-ACTIN (Thermo Fisher Scientific, Waltham,MA). For animal experiments, eye tissues were harvested from E15.5 or E18.5 mouse embryos or 8-month-old mice. The lens, cornea, and optic nerve were dissected, RNA was isolated and cDNA generated, and TaqMan Assays conducted with inventoried probes for Myrf and Tmem98 (Thermo Fisher Scientific, Waltham, MA). Critical cycle threshold levels of each sample were normalized to levels of β-actin for human cells and Hprt for animal experiments. Fold activity was calculated using the ΔΔCt method and reported relative to controls.
In silico modeling. PONDR (23) and AlphaFold2 (22) were used for in silico modeling of the WT-MYRF and dG-MYRF conserved C2 domain amino acid sequence. PONDR was used to predict differences in intrinsically ordered and disordered regions. VXLT combines 3 previous neural network models (VL1, XN, and XC) trained on NMR and x-ray crystallography data with varying lengths of disordered regions. VSL2 combines 2 predictors optimized for long (>30 residues) and short (≤30 residues) disordered regions (20, 21, 23, 65). Alphafold2 was used for de novo prediction of 3D structure of the C2 domain in the presence or absence of the C-terminal frameshift variant amino acid sequence.
Generation of humanized Myrf alleles in mice. Mice were genetically engineered using CRISPR-Cas9 to replace mouse Myrf exon 26 with the homologous human MYRF exon 27 and fused it with mouse exon 25, including the C-terminal coding region the human-3′UTR sequence (Supplemental Figure 3). This humanized Myrf allele (MyrfhumWT) was generated by injecting fertilized eggs from C57BL/6J matings with a pair of spCas9 guide RNAs (25 ng/μL each) (gRNA, protospacer adjacent motif=NGG) that are 253 bps apart in outward orientation together with the spCas9 protein (NEB, 30 ng/μL). Sequences of the gRNAs are listed below: the upstream gRNA 5′-GCTCACCAGCAATGTCACC-3′ and the downstream gRNA 5′-GGCAGTGCCACAGCCGGGAC-3′. A single stranded oligo of 327 bps (IDT Megamer) carrying the human Exon27 sequence and homology arms of 100 bp each is also included in the injection mixture as the repair template for single strand DNA mediated homologous recombination (HR). The sequence of this HR template is 5′- TAGGACAAGGCTGCCCTGACCTCTGTCTTCCCTGTCTCTCTAGGGCACC TCTCATCAGTGGCCAGTAACCATCCTGTCCTTCCGTGAATTCACATACCACTTtCGGGTGACATTGCTGg GTCAGGCCAACTGCAGTTCAGAGGCTCTCGCCCAGCCAGCCACAGACTACCACTTCCACTTCTACCGCCTGTG TGACTGAGCTGCCCTCCTGACAGTGCCACAGCCGcGACTGGAGTCCCTGGGCCCTCAACACTGGATGCA AATGTGTTACACTGGAGCCTGCTGCAGGCCAGCTCTCTGCTCTTCACTGCTTCCCTTGACTGGGGA-3′ (327 bps). Founder mice were genotyped using primers (Forward: 5′-GTGG CTACCCTTGCTCTCAA-3′, Reverse: 5′-GGCGAGAGCCTCTGAACTG-3′).
We generated a matched MyrfhumdG mouse that recapitulates the splice site mutation found in our previously described large familial cohort with nanophthalmos, starting with heterozygous MyrfhumWT/+, and using the following guide RNA (5′-CACTTCCGGGTGACATTGCT-3′) and ssDNA repair template: 5′- TGGCCAGTAACCATCCTGTCCTTCCGTGAATTCACATACXCACTTCCGGGTGA CATTGCTGGTCAGGCCAACTGCAGTTCAGAGGCTCTCGCCCAG CCAGCCACAGACTACCACTTCCACT-3′. RxCre;Myrffl/fl mice have been previously described (8). Mice were genotyped using standard methods (66) and primers listed above, followed by restriction digest with the BseYI restriction enzyme site to distinguish between the MyrfhumdG and MyrfhumWT alleles. Animals were also initially genotyped for rd1, rd8, and rd10 according to established methods, to eliminate retinal degeneration alleles common in C57BL/6J mouse strains (67).
RNAscope in situ hybridization. Heads were harvested from E15.5 embryos. Samples were fixed in 4% buffered PFA (0.1M NaPO4, pH 7.3) overnight at 4°C, dehydrated through increasing concentrations of ethanol up to 70%. Samples were dehydrated, processed and sectioned as previously described (8, 10). RNAscope was performed using reagents from the RNAscope Multiplex Fluorescent Detect V2 kit (Advanced Cell Diagnostics [ACD], Newark, CA, #323110) as previously described (8, 10).
Retinal layer measurements, whole eye measurements, and cell counting. Eyes from 12-month animals were enucleated and aligned to capture optic nerve and central cornea in the same plane, and images were captured on the Leica MX10F dissecting microscope (Leica, Wetzlar, Germany). Ocular axial length was measured at 12 months using ImageJ (NIH) 1.51m9 software (68), by marking a line from the central cornea to the base of the optic nerve. For the analysis, the following number of animals/eyes were used: MyrfhumWT/+ control, n = 4 animals, 8 eyes; MyrfhumdG/+, n = 5 animals, 9 eyes.
Mouse ocular imaging and electrophysiology. Eyes from 3-, 6-, 9-, and 12-month old control (MyrfhumWT/+; n = 8 eyes, 4 mice) and heterozygous mutant mice (MyrfhumdG/+; n = 9 eyes, 10 mice) were evaluated sequentially in vivo by fundus photography, spectral domain OCT, and ERG as previously described (8). Data from MyrfhumdG/+ and controls were evaluated for statistical significance with 1-way ANOVA in GraphPad Prism 10 (GraphPad Software, San Diego, CA, USA). If significant, subsequent pairwise comparisons were done with 2-tailed Student’s t test. A-wave and B-wave amplitudes from MyrfhumdG/+ eyes and control littermates were evaluated using 2-tailed Student’s t test in GraphPad Prism 10.
RPE flat mounts. Eyes were enucleated from 12-month mice, and cornea, lens, optic nerve, and retina tissues were removed. At least 3 animals/eyes per genotype/age were used. Optic cups were fixed, washed, and blocked as previously described (8). Primary antibody incubation occurred overnight at 4°C: TMEM98 (1:500, 14731-1-AP, Proteintech, Rosemont, IL), Phalloidin (1:400, #A12380, Thermo Fisher, Scientific, Waltham, MA). RPE samples were washed in PBS, incubated with Alexa-conjugated fluorescent secondary antibodies, and mounted as previously described (8). Images were taken on a Lecia DM6000 compound microscope.
REShAPE RPE morphometric analysis. RPE flat mounts (see above) were prepared from 12-month-old control (MyrfhumWT/humWT; n = 4 eyes, 4 mice), heterozygous (MyrfhumWT/humdG; n = 5 eyes, 10 mice) from a single cohort. RPE flat mounts were stained as above with phalloidin to outline RPE cell borders. They were imaged using the LASX Navigator tiling software on a Lecia DM6000 compound microscope at 20X. Tiled images were fed into the REShAPE software (27), processed, and analyzed by the REShAPE software to look at morphometrics such as cell size, aspect ratio, hexagonality, and number of neighboring cells. Data points for RPE morphometrics from each tile were combined per biological replicate. Control and heterozygous RPE morphometrics were analyzed by performing a Student’s t test on the median value.
scRNA-seq analysis. Eyes from E17.5 MyrfhumWT/humWT (n = 3 mice, 6 eyes) and MyrfhumdG/humdG (n = 3 mice, 6 eyes) were dissected removing the lens and optic nerve and leaving only the optic cup (retina/RPE/sclera/cornea) and dissociated as previously described (10). Eyes were processed for scRNA-seq at the University of Michigan Advanced Genomics Core as previously described using the CellRanger 7.1.0 pipeline (10, 69). SEURAT 3.1.2 package (70, 71) was used to performed integration, unsupervised clustering, and cell type identification on scRNA-seq of E17.5 optic cups from this study with previously published scRNA-seq datasets from E15.5 and P0 RxCre;Myrffl/fl mouse optic cups and paired controls (10). Quality control thresholds were set to nFeature_RNA > 200 and percent.mt < 15. SEURAT 5.1.0 package was used to generate up- and downregulated differentially expressed genes in the RPE cluster. The ClusterProfiler 4.14.6 (72) package was used to look for enriched GO terms and GSEA (73). UCell was used to assign “scores” to enriched GO term and GSEA pathways in MyrfhumWT/humWT (n = 3 mice, 6 eyes) and MyrfhumdG/humdG mice on a per cell basis (74).
Patient collection, sequencing, and analysis. The patient cohort consisting of individuals with axial length < 21 mm or high hyperopia > +5.50 spherical equivalent was previously described (34). Clinical and biometric data were reviewed for patients. DNA from whole blood or saliva was extracted according to standard procedures as before. Patient samples were sequenced at the National Intramural Sequencing Center (Rockville, MD) by Illumina short-read WGS using the Illumina NovaSeq platform (Illumina, San Diego, CA). Reads were aligned and variants were called and annotated via a customized pipeline (https://github.com/Bin-Guan/NGS_genotype_calling; commit ID 32a0371, and https://github.com/Bin-Guan/variant_prioritization; commit ID 5cfb4f8).
Minigene splicing assays and patient blood RNA analysis. P04825 and P01965 WT and variant sequences were cloned into the pSLP3 exon trap vector. HEK293T (ATCC, #CRL-2316, American Type Culture Collection Manassas, VA) cells were seeded in 6-well plates at a density of 75,000 cells per well. Plasmid DNA (970 ng) and 30 ng GFP plasmid were transfected into HEK293T cells using FuGENE 6 transfection reagent (Promega, Madison, WI) at a 3:1 ratio. Cells were incubated overnight and then harvested the next day for RNA isolation using the QIAGEN Mini RNA Isolation Kit (Qiagen, Germantown, MD). cDNA was generated using SuperScript III and random primers following manufacturer’s protocols. Splicing within the exon trap vector was analyzed by amplifying across the exon trap with for V1 primer 5′-TCTGAGTCACCTGGACAACC-3′ and rev V2 primer 5′-ATCTCAGTGGTATTTGTGAGC-3′. PCR products were confirmed by Sanger sequencing. The PCR products were cloned into the TOPO TA cloning vector (Thermo Fisher Scientific, Waltham, MA). Eighteen colonies were picked and sequenced to test the frequency of splicing into the cryptic splice acceptor site.
Blood was collected from subject P04818 and stored in PAXgene tubes at –80°C until RNA preparation (Qiagen, Germantown, MD). Prior to isolating the RNA, the blood sample was equilibrated to RT for 2 hours. RNA was isolated using the protocol provided with the PAXgene Blood RNA Kit, PreAnalytix (Qiagen, Germantown, MD). RNA was treated with DNaseI (Invitrogen, Carlsbad, CA) and transcribed into cDNA using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems Foster City, CA). cDNA was amplified using a forward oligo (MYRF exon 23; 5′-ATCACCTCCCAGTACTGTGC-3′) and a reverse oligo (MYRF exon 25; 5′-GGTGTGGAGACTCTGTGGAA-3′) with standard PCR cycling and 55°C annealing. Normal splicing was represented by a 203 bp band and splicing including the pseudoexon was represented by a 324 bp band. A gel fragment corresponding to 150–350 bp was isolated from the amplified patient cDNA and control. DNA was isolated using the QIAEX gel extraction kit (Qiagen, Germantown, MD) and cloned into a TA vector using the TOPO TA Cloning for Sequencing Kit (Invitrogen, Carlsbad, CA). Twenty colonies were collected and analyzed by PCR and sequenced to calculate the occurrence of splicing into the cryptic splice site.
Statistics. A χ2 test was used to detect significant skewing of expected Mendelian ratios of genotypes at E12.5–P0. A Wilcoxon Rank Sum test was used to identify DEGs between 2 groups of cells in the scRNA-seq dataset. For DEG analysis, an Padj value was calculated based on Bonferroni correction using all genes in the dataset. DEGs were considered significant with a P-adjusted value less than 0.1. One-way ANOVA was used when comparing the means of 3 or more groups. A 2-way ANOVA with multiple comparisons was used when comparing the means of 3 or more groups across genotypes and timepoints. A Student’s t test was used when comparing the means of only 2 groups. Unless otherwise noted, in the figure legend data represent mean ± SD.
Study approval. Patient work was approved by the IRB at the University of Michigan under protocol no. HUM00046159. All patients provided written informed consent for their inclusion. Mouse studies were approved by the IACUC at the University of Michigan under protocol nos. PRO00010115 and PRO00011748. This study was carried out in accordance with the Tenets of the Declaration of Helsinki.
Data availability. scRNA-seq data are available at the Gene Expression Omnibus (GEO, National Center for Biotechnology Information, National Library of Medicine; GSE310659). Genomic data are accessible at dbGAP (accession no. phs004462.v1.p1). The code used to process scRNA-sequencing data as well as analyze REShAPE morphometric and scRNA-sequencing differential gene expression has been uploaded as supplemental.txt files. Primary data are provided for all the experiments in the Supporting Data Values file. All additional relevant data can be found within the article, and its supplementary information and primary data are available upon request from the corresponding author.
-
Author contributions
GMR and LP wrote the manuscript. GMR precedes MLB in co–first authors as she took the lead in the project and wrote the manuscript. GMR, MLB, and LP designed research studies, conducted experiments, acquired data, and analyzed data. SQW, CT, NTY, DK, RSL, and DS conducted experiments, acquired data, and analyzed data. LP, QZ, JYSH, JMLM, LD, SEM, JER, and RBH provided reagents. BG and DFH analyzed data.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- NEI K08 EY032098 (to LP).
- E. Matilda Ziegler Foundation for the Blind (to LP).
- Bright Focus Foundation (M2022011N) (to LP).
- Research to Prevent Blindness Career Development Award (to LP).
- Knight Templar Eye Foundation funding (to LP).
- Vision Research Core (NEI P30-EY007003) (to LP).
- Shared Instrument Grant S10 (NIH SIG grant S10OD28612-01-A1) (to LP).
- NEI T32 EY013934 (to GMR).
- James Grosfeld Initiative for Dry AMD Trainee Grant (to GMR).
- NEI intramural funding (RBH, BG, and LD).
- NEI K08 - EY033420 (to JMLM).
- The James Grosfeld Initiative for Dry AMD (to JMLM.)
- Discovering Hope Foundation funding (to JMLM).
- NEI R01 EY011671 (to JER).
- BrightFocus Foundation Postdoctoral Fellowship M2024001F (to JYSH).
- Michigan Pioneer Fellows Program (to JYSH).
- Intramural Research Program of the National Institutes of Health (NIH), The National Eye Institute.
-
Acknowledgments
We would like to thank the Thom Saunders, Elizabeth Hughes, and the University of Michigan Transgenic Core for helping to generate our humanized mouse model, the Besirli lab (Eric Weh) for help with REShAPE software, the Vision Research Core (Chengmao Lin and Sarah Sheskey) for technical assistance with fundus imaging and OCT analysis, and other members of the Miller lab (Gillian Gulette) for their contribution of human primary cells, protocols, and expertise. We thank Enrique Rodriguez-Boulan lab for the RPE bleaching protocol, the Advanced Genomics Core for assistance with scRNA-seq, Ben Emery for helpful advice, and Sally Camper, Andrew Lieberman, Tom Wilson for helpful discussions. No federal funds were used for HFT research. The contributions of the NIH authors were made as part of their official duties as NIH federal employees, in compliance with agency policy requirements, and considered Works of the United States Government. However, the findings and conclusions presented in this paper are those of the authors and do not necessarily reflect the views of the NIH or the U.S. Department of Health and Human Services.
Author for correspondence: Lev Prasov, KEC 229, 1000 Wall St., Ann Arbor, Michigan 48105, USA. Phone: 734.647.4447; Email: lprasov@umich.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2026, Rozumek et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(6):e194681.https://doi.org/10.1172/jci.insight.194681.
-
References
- Bujalka H, et al. MYRF is a membrane-associated transcription factor that autoproteolytically cleaves to directly activate myelin genes. PLoS Biol. 2013;11(8):e1001625.
- Li Z, et al. A Bacteriophage tailspike domain promotes self-cleavage of a human membrane-bound transcription factor, the myelin regulatory factor MYRF. PLoS Biol. 2013;11(8):e1001624.
- Kaplan JD, et al. MYRF-Related Cardiac Urogenital Syndrome. In: Adam MP, et al., eds. GeneReviews. University of Washington; 2022.
- Qi H, et al. De novo variants in congenital diaphragmatic hernia identify MYRF as a new syndrome and reveal genetic overlaps with other developmental disorders. PLoS Genet. 2018;14(12):e1007822.
- Pinz H, et al. De novo variants in Myelin regulatory factor (MYRF) as candidates of a new syndrome of cardiac and urogenital anomalies. Am J Med Genet A. 2018;176(4):969–972.
- Rossetti LZ, et al. Review of the phenotypic spectrum associated with haploinsufficiency of MYRF. Am J Med Genet A. 2019;179(7):1376–1382.
- Hamanaka K, et al. MYRF haploinsufficiency causes 46,XY and 46,XX disorders of sex development: bioinformatics consideration. Hum Mol Genet. 2019;28(14):2319–2329.
- Garnai SJ, et al. Variants in myelin regulatory factor (MYRF) cause autosomal dominant and syndromic nanophthalmos in humans and retinal degeneration in mice. PLoS Genet. 2019;15(5):e1008130.
- Ouyang J, et al. Truncation mutations in MYRF underlie primary angle closure glaucoma. Hum Genet. 2023;142(1):103–123.
- Brinkmeier ML, et al. Myelin regulatory factor (MYRF) is a critical early regulator of retinal pigment epithelial development. PLoS Genet. 2025;21(4):e1011670.
- Yu X, et al. Myelin regulatory factor deficiency is associated with the retinal photoreceptor defects in mice. Vis Neurosci. 2021;38:E005.
- Yu X, et al. Nanophthalmos-associated MYRF gene mutation causes ciliary zonule defects in mice. Invest Ophthalmol Vis Sci. 2021;62(3):1.
- Qi YB, et al. MYRF: a unique transmembrane transcription factor- from proteolytic self-processing to its multifaceted roles in animal development. Bioessays. 2024;46(4):e2300209.
- An H, et al. Functional mechanism and pathogenic potential of MYRF ICA domain mutations implicated in birth defects. Sci Rep. 2020;10(1):814.
- Fan C, et al. Functional mechanisms of MYRF DNA-binding domain mutations implicated in birth defects. J Biol Chem. 2021;296:100612.
- Tao J, et al. MFRP, PRSS56, and MYRF account for 60.5% of a Chinese cohort with nanophthalmos. Clin Exp Ophthalmol. 2025;53(2):194–208.
- Guo C, et al. Detection of clinically relevant genetic variants in chinese patients with nanophthalmos by trio-based whole-genome sequencing study. Invest Ophthalmol Vis Sci. 2019;60(8):2904–2913.
- Kim D, et al. Homo-trimerization is essential for the transcription factor function of Myrf for oligodendrocyte differentiation. Nucleic Acids Res. 2017;45(9):5112–5125.
- Schulz EC, et al. Crystal structure of an intramolecular chaperone mediating triple-beta-helix folding. Nat Struct Mol Biol. 2010;17(2):210–215.
- Garner E, et al. Predicting binding regions within disordered proteins. Genome Inform Ser Workshop Genome Inform. 1999;10:41–50.View this article via: PubMed Google Scholar
- Dunker AK, et al. Intrinsically disordered protein. J Mol Graph Model. 2001;19(1):26–59.
- Jumper J, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596(7873):583–589.
- PONDR. Predictor of Natural Disordered Regions (PONDR). https://pondr.com/ Accessed February 10, 2026.
- Zhang Q, et al. A platform for assessing outer segment fate in primary human fetal RPE cultures. Exp Eye Res. 2019;178:212–222.
- Miller JML, et al. Regression of drusen or vitelliform material heralding geographic atrophy: correlation between clinical observations and basic science. Graefes Arch Clin Exp Ophthalmol. 2021;259(7):2051–2053.
- Cross SH, et al. The nanophthalmos protein TMEM98 inhibits MYRF self-cleavage and is required for eye size specification. PLoS Genet. 2020;16(4):e1008583.
- Hotaling N, et al. REShAPE – a cloud-computing based cell morphometry analyzer for development of a release criteria of clinical grade retinal pigment epithelium. Invest Ophthalmol Vis Sci. 2017;58(8):396.
- Campochiaro PA, et al. The extracellular matrix of human retinal pigment epithelial cells in vivo and its synthesis in vitro. Invest Ophthalmol Vis Sci. 1986;27(11):1615–1621.View this article via: PubMed Google Scholar
- Singh K, et al. Cigarette smoke and biological age induce degenerative heterogeneity in retinal pigment epithelium [preprint]. https://doi.org/10.1101/2025.01.27.635096 Posted on bioRxiv January 28, 2025.
- Dale D, Summers JA. Bidirectional regulation of SERPINE3 gene expression in the RPE during visually induced changes in ocular growth. Invest Ophthalmol Vi Sci. 2024;65(7):1180.
- Collery RF. Expression of SERPINE3 is bidirectionally altered in refractive error. Invest Ophthalmol Vis Sci. 2023;64(8):859.
- Zhang X, et al. KCNQ5/Kv7.5 potassium channel expression and subcellular localization in primate retinal pigment epithelium and neural retina. Am J Physiol Cell Physiol. 2011;301(5):C1017–C1026.
- Zhang X, Hughes BA. KCNQ and KCNE potassium channel subunit expression in bovine retinal pigment epithelium. Exp Eye Res. 2013;116:424–432.
- Prasov L, et al. Novel TMEM98, MFRP, PRSS56 variants in a large United States high hyperopia and nanophthalmos cohort. Sci Rep. 2020;10(1):19986.
- Jaganathan K, et al. Predicting splicing from primary sequence with deep learning. Cell. 2019;176(3):535–548.
- Emery B, et al. Myelin gene regulatory factor is a critical transcriptional regulator required for CNS myelination. Cell. 2009;138(1):172–185.
- Xu Q, et al. Comprehensive genetic landscapes and clinical heterogeneity in nanophthalmos: new insights from a large Chinese cohort. [published online September 19, 2025]. J Med Genet. https://doi.org/10.1136/jmg-2025-110905. View this article via: PubMed Google Scholar
- Siggs OM, et al. The genetic and clinical landscape of nanophthalmos and posterior microphthalmos in an Australian cohort. Clin Genet. 2020;97(5):764–769.
- Koli S, et al. Identification of MFRP and the secreted serine proteases PRSS56 and ADAMTS19 as part of a molecular network involved in ocular growth regulation. PLoS Genet. 2021;17(3):e1009458.
- Carricondo PC, et al. Nanophthalmos: a review of the clinical spectrum and genetics. J Ophthalmol. 2018;2018:2735465.
- Nair S, et al. Retinal PRSS56: a critical GO signal in axial elongation and myopia. Invest Ophthalmol Vis Sci. 2025;66(8):4574.
- Zenteno JC, et al. Homozygosity mapping identifies the Crumbs homologue 1 (Crb1) gene as responsible for a recessive syndrome of retinitis pigmentosa and nanophthalmos. Am J Med Genet A. 2011;155A(5):1001–1006.
- Boon CJF, et al. The spectrum of ocular phenotypes caused by mutations in the BEST1 gene. Prog Retin Eye Res. 2009;28(3):187–205.
- Abouzeid H, et al. A G1103R mutation in CRB1 is co-inherited with high hyperopia and Leber congenital amaurosis. Ophthalmic Genet. 2006;27(1):15–20.
- Cross SH, et al. Missense mutations in the human nanophthalmos gene TMEM98 cause retinal defects in the mouse. Invest Ophthalmol Vis Sci. 2019;60(8):2875–2887.
- Hassall MM, et al. Phenotypic consequences of a nanophthalmos-associated TMEM98 variant in human and mouse. Sci Rep. 2023;13(1):11017.
- Tkatchenko TV, et al. Analysis of postnatal eye development in the mouse with high-resolution small animal magnetic resonance imaging. Invest Ophthalmol Vis Sci. 2010;51(1):21–27.
- Zhou X, et al. The development of the refractive status and ocular growth in C57BL/6 mice. Invest Ophthalmol Vis Sci. 2008;49(12):5208–5214.
- Paritala A, et al. Correlation of vitreous chamber depth with ocular biometry in high axial myopia. Indian J Ophthalmol. 2022;70(3):914–920.
- Takkar B, et al. Evaluation of the vitreous chamber depth: an assessment of correlation with ocular biometrics. Indian J Ophthalmol. 2019;67(10):1645–1649.
- Fan Q, et al. Meta-analysis of gene-environment-wide association scans accounting for education level identifies additional loci for refractive error. Nat Commun. 2016;7(1):11008.
- Fuse N, et al. Genome-wide association study of axial length in population-based cohorts in Japan: the Tohoku medical megabank organization eye study. Ophthalmol Sci. 2022;2(1):100113.
- Tedja MS, et al. Genome-wide association meta-analysis highlights light-induced signaling as a driver for refractive error. Nat Genet. 2018;50(6):834–848.
- Milan M, et al. Pancreatic cancer cells require the transcription factor MYRF to maintain ER homeostasis. Dev Cell. 2020;55(4):398–412.
- Gupta N, et al. Diverse clinical manifestations and intrafamilial variability due to an inherited recurrent MYRF variant. Am J Med Genet A. 2022;188(7):2187–2191.
- Ament IH, et al. Long-read RNA sequencing: a transformative technology for exploring transcriptome complexity in human diseases. Mol Ther. 2025;33(3):883–894.
- Marx V. Method of the year: long-read sequencing. Nat Methods. 2023;20(1):6–11.
- Sage J, et al. Acute mutation of retinoblastoma gene function is sufficient for cell cycle re-entry. Nature. 2003;424(6945):223–228.
- Lynn SA, et al. A convenient protocol for establishing a human cell culture model of the outer retina. F1000Res. 2018;7:1107.
- Fronk AH, Vargis E. Methods for culturing retinal pigment epithelial cells: a review of current protocols and future recommendations. J Tissue Eng. 2016;7:2041731416650838.
- Maminishkis A, et al. Confluent monolayers of cultured human fetal retinal pigment epithelium exhibit morphology and physiology of native tissue. Invest Ophthalmol Vis Sci. 2006;47(8):3612–3624.
- Hass DT, et al. Medium depth influences O2 availability and metabolism in human RPE cultures. Invest Ophthalmol Vis Sci. 2023;64(14):4.
- Sharma R, et al. Triphasic developmentally guided protocol to generate retinal pigment epithelium from induced pluripotent stem cells. STAR Protoc. 2022;3(3):101582.
- Fernandes M, et al. Stem cell-derived retinal pigment epithelial layer model from adult human globes donated for corneal transplants. Curr Protoc Stem Cell Biol. 2018;45(1):e53.
- PONDR. Predictor of Naturally Disordered Regions - disorder protein predictor. https://pondr.com/pondr-tut2.html Accessed February 10, 2026.
- Miller SA, et al. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988;16(3):1215.
- Chang B, et al. Survey of common eye diseases in laboratory mouse strains. Invest Ophthalmol Vis Sci. 2013;54(7):4974–4981.
- Schneider CA, et al. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9(7):671–675.
- Zheng GXY, et al. Massively parallel digital transcriptional profiling of single cells. Nat Commun. 2017;8(1):14049.
- Hao Y, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184(13):3573–3587.
- Stuart T, et al. Comprehensive integration of single-cell data. Cell. 2019;177(7):1888–1902.
- Yu G, et al. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–287.
- Subramanian A, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–15550.
- Andreatta M, Carmona SJ. UCell: robust and scalable single-cell gene signature scoring. Comput Struct Biotechnol J. 2021;19:3796–3798.
-
Version history
- Version 1 (February 26, 2026): In-Press Preview
- Version 2 (March 23, 2026): Electronic publication